Featured Products
Your Dynamic Snippet will be displayed here... This message is displayed because you did not provided both a filter and a template to use.

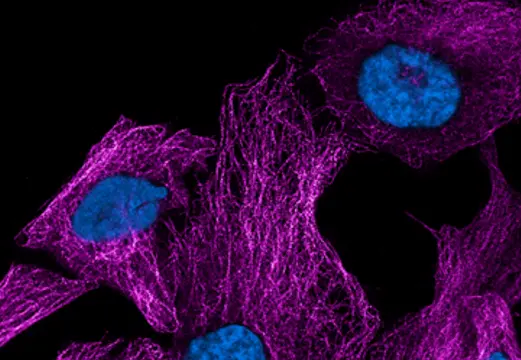
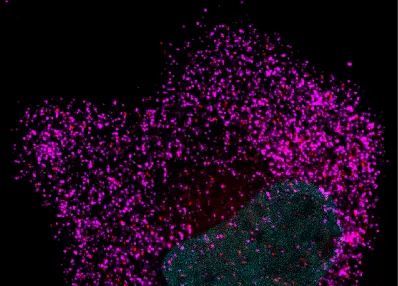
Picture of the Month
Show Us What You See!
Submit your most stunning Expansion Microscopy image for a chance to be featured and win Amazon vouchers. Share your science, inspire others, and showcase the power of expanded imaging.